silx #
silx provides applications and Python modules to support the development of data assessment, reduction, and analysis at synchrotron radiation facilities. It provides reading/writing tools for different file formats, data reduction routines, and a set of Qt widgets to browse and visualise data.
Installation#
You can install silx via pip, conda, or apt on Debian-flavoured Linux distributions, using the commands listed for each case. Self contained macOS applications for Intel and Apple Silicon and a Windows installer are also available for download (see the links below).
pip install silx[full]
conda install -c conda-forge silx
sudo apt install silx
Installer (.exe) |
Archive (.zip) |
|---|---|
Intel (x86_64) |
Apple Silicon (arm64) |
|---|---|
Applications#
silx view
Unified viewer supporting HDF5, SPEC and image file formats
silx compare
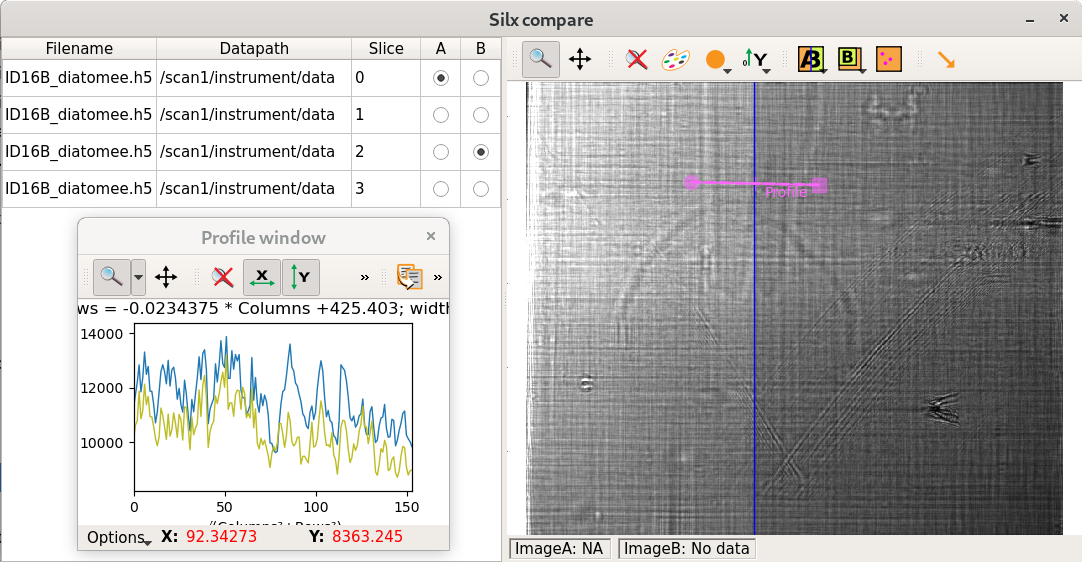
User interface to compare 2D data from files
silx convert
Converter of legacy file formats into HDF5 file
Python modules#
silx.gui
1D and 2D visualization widgets and associated tools
OpenGL-based 3D visualization widgets
A unified HDF5, SPEC and image data file browser and n-dimensional dataset viewer
silx.opencl
Image alignment (SIFT)
Image processing (median filter, histogram)
Filtered backprojection for tomography
silx.io
silx.math
